SQL Quick Start Using a FLOAT32 Vector Generator
A PL/SQL program that creates a vector generator is included along with example queries and results, providing a simple way to get started with Oracle AI Vector Search without a vector embedding model.
genvec that you can then use, for example,
to experiment with similarity searches.
The following instructions assume you already have access to a database account with
sufficient privileges (minimally the DB_DEVELOPER_ROLE role).
Note:
Do not use the vector generator on production databases. The program is made available for testing and demo purposes.- Create the
genvectable.DROP TABLE genvec PURGE; CREATE TABLE genvec ( id number, -- id of the generated vector v VECTOR, -- generated vector name VARCHAR2(500), -- name for the generated vector: C1 to Cn are centroids, Cx_y is vector number y in cluster number x nv VECTOR, -- normalized version of the generated vector ly number -- random number you can use to filter out rows in addition to similarity search on vectors ); - Create the package
vector_gen_pkgand its associated package body.Here you create the vector generator package:
CREATE OR REPLACE PACKAGE vector_gen_pkg AS TYPE t_vectors IS TABLE OF vector INDEX BY PLS_INTEGER; FUNCTION get_coordinate( input_string CLOB, i PLS_INTEGER ) RETURN NUMBER; PROCEDURE generate_vectors( num_vectors IN PLS_INTEGER, -- Number of vectors to generate dimensions IN PLS_INTEGER, -- Number of dimensions of each vector num_clusters IN PLS_INTEGER, -- Number of clusters to create cluster_spread IN NUMBER, -- Relative closeness of each vector in each cluster (using standard deviation) min_value IN NUMBER, -- Minimum value for a vector coordinate max_value IN NUMBER -- Maximum value for a vector coordinate ); END vector_gen_pkg; /And the package body:
CREATE OR REPLACE PACKAGE BODY vector_gen_pkg AS ------------------------------------------- ------------------------------------------- ---- V E C T O R G E N E R A T O R ----- ------------------------------------------- ------------------------------------------- -- Version 1.0 --------- ------------------------------------------- ------------------------------------------- ---- DO NOT USE ON PRODUCTION DATABASES --- ---- ONLY FOR TESTING AND DEMO PURPOSES --- ------------------------------------------- FUNCTION get_coordinate( input_string CLOB, i PLS_INTEGER ) RETURN NUMBER IS start_pos NUMBER; end_pos NUMBER; comma_pos NUMBER; coord VARCHAR2(100); comma_count NUMBER := 0; commas NUMBER; working_string CLOB; BEGIN -- Remove leading and trailing brackets working_string := input_string; working_string := TRIM(BOTH '[]' FROM working_string); commas := LENGTH(working_string) - LENGTH(REPLACE(working_string, ',', '')); -- Initialize positions start_pos := 1; end_pos := INSTR(working_string, ',', start_pos); IF i<=0 OR i>commas+1 THEN RETURN NULL; END IF; -- Loop through the string to find the i-th coordinate LOOP IF comma_count + 1 = i THEN IF end_pos = 0 THEN -- If there's no more comma, the coordinate is the rest of the string coord := SUBSTR(working_string, start_pos); ELSE coord := SUBSTR(working_string, start_pos, end_pos - start_pos); END IF; RETURN coord; END IF; -- Move to the next coordinate comma_count := comma_count + 1; start_pos := end_pos + 1; end_pos := INSTR(working_string, ',', start_pos); -- Exit loop if no more coordinates EXIT WHEN start_pos > LENGTH(working_string); END LOOP; -- If the function hasn't returned yet, the index was out of bounds RETURN NULL; END; PROCEDURE generate_random_vector( dimensions IN PLS_INTEGER, min_value IN NUMBER, max_value IN NUMBER, vec OUT vector ) IS e CLOB; BEGIN e := '['; FOR i IN 1..dimensions-1 LOOP e := e || DBMS_RANDOM.VALUE(min_value, max_value) ||','; END LOOP; e := e || DBMS_RANDOM.VALUE(min_value, max_value) ||']'; vec := VECTOR(e); END generate_random_vector; PROCEDURE generate_clustered_vector( centroid IN vector, cluster_spread IN NUMBER, vec OUT vector ) IS e CLOB; d number; BEGIN d := VECTOR_DIMENSION_COUNT(centroid); e := '['; FOR i IN 1 .. d-1 LOOP e := e || (get_coordinate(to_clob(centroid),i) + (DBMS_RANDOM.NORMAL * cluster_spread)) ||','; END LOOP; e := e || (get_coordinate(to_clob(centroid),VECTOR_DIMENSION_COUNT(centroid)) + (DBMS_RANDOM.NORMAL * cluster_spread)) || ']'; vec := VECTOR(e); END generate_clustered_vector; FUNCTION normalize_vector(vec IN vector) RETURN vector IS e CLOB; v CLOB; n number; d number; BEGIN n := VECTOR_NORM(vec); v := to_clob(vec); d := VECTOR_DIMENSION_COUNT(vec); e := '['; FOR i IN 1 .. d-1 LOOP e := e || (get_coordinate(v,i)/n) ||','; END LOOP; e := e || (get_coordinate(v,d)/n) || ']'; RETURN VECTOR(e); END normalize_vector; PROCEDURE generate_vectors( num_vectors IN PLS_INTEGER, -- Must be 1 or above dimensions IN PLS_INTEGER, -- Must be above 1 but less than 500 num_clusters IN PLS_INTEGER, -- Must be 1 or above cluster_spread IN NUMBER, -- Must be grather than 0 min_value IN NUMBER, max_value IN NUMBER ) IS centroids t_vectors; vectors_per_cluster PLS_INTEGER; remaining_vectors PLS_INTEGER; vec vector; idx PLS_INTEGER := 1; max_id NUMBER; working_vector VECTOR; BEGIN IF (num_vectors) <=0 OR (num_clusters < 1) OR (num_vectors < num_clusters) OR (dimensions <= 0) OR (dimensions > 500) OR (cluster_spread <= 0) OR (min_value >= max_value) THEN RETURN; END IF; SELECT MAX(id) INTO max_id FROM genvec; IF max_id IS NULL THEN max_id := 0; END IF; -- Generate cluster centroids FOR i IN 1..num_clusters LOOP generate_random_vector(dimensions, min_value, max_value, centroids(i)); working_vector := normalize_vector(centroids(i)); INSERT INTO genvec VALUES (max_id + idx, centroids(i), 'C'||i, working_vector, DBMS_RANDOM.VALUE(3,600000000)); idx := idx + 1; END LOOP; -- Calculate vectors per cluster vectors_per_cluster := TRUNC(num_vectors / num_clusters); remaining_vectors := num_vectors MOD num_clusters; -- Generate vectors for each cluster IF vectors_per_cluster > 1 THEN FOR i IN 1..num_clusters LOOP FOR j IN 1..(vectors_per_cluster - 1) LOOP generate_clustered_vector(centroids(i), cluster_spread, vec); working_vector := normalize_vector(vec); INSERT INTO genvec VALUES (max_id + idx, vec, 'C'||i||'-'||j, working_vector, DBMS_RANDOM.VALUE(3,600000000)); idx := idx + 1; END LOOP; END LOOP; END IF; -- Handle remaining vectors: all associated with cluster 1 IF remaining_vectors > 0 THEN FOR j IN 1..remaining_vectors LOOP generate_clustered_vector(centroids(1), cluster_spread, vec); working_vector := normalize_vector(vec); INSERT INTO genvec VALUES (max_id + idx, vec, 'C1-'||idx, working_vector, DBMS_RANDOM.VALUE(3,600000000)); idx := idx + 1; END LOOP; END IF; COMMIT; END generate_vectors; END vector_gen_pkg; / - After you have your vector generator set up, you can run this and the following steps
to understand how to use it.
Run the
generate_vectorsprocedure of thevector_gen_pkgpackage with sample values:BEGIN vector_gen_pkg.generate_vectors( num_vectors => 100, -- Number of vectors to generate. Must be 1 or above dimensions => 3, -- Number of dimensions of each vector. Must be above 1 but less than 500 num_clusters => 6, -- Number of clusters to create. Must be 1 or above cluster_spread => 1, -- Relative closeness of each vector in each cluster (using standard deviation). Must be grather than 0 min_value => 0, -- Minimum value for a vector coordinate max_value => 100 -- Maximum value for a vector coordinate. Min value must be smaller than max value ); END; / - Run a
SELECTstatement to view the newly generated vectors.SELECT name, v FROM genvec;Example output:
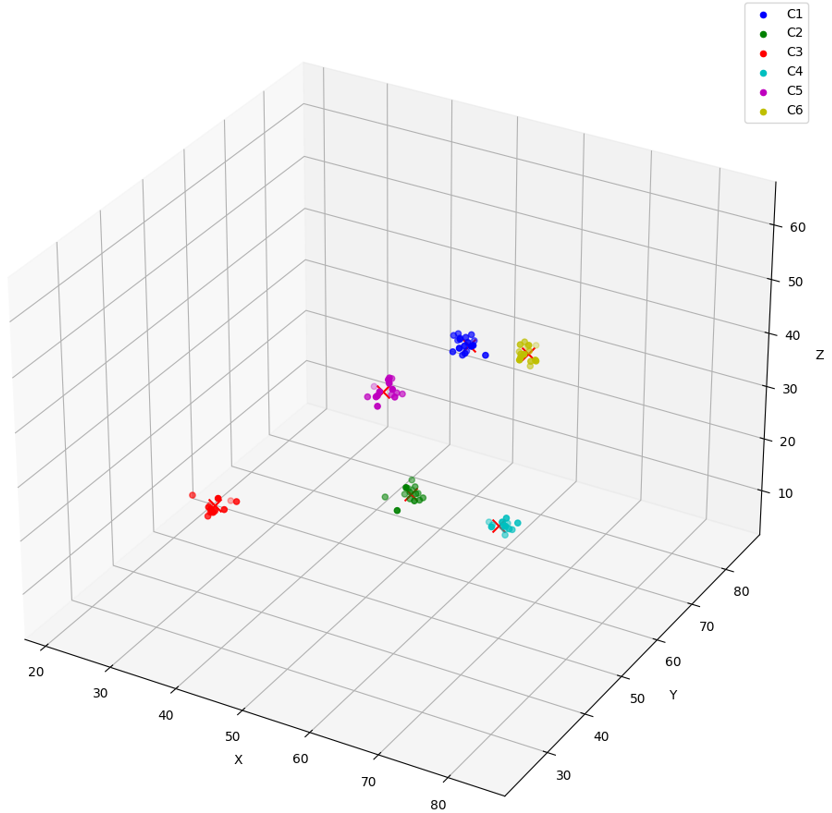
NAME V _______ ____________________________________________________ C1 [6.35792809E+001,5.28954163E+001,5.16500435E+001] C2 [5.67991257E+001,5.00640755E+001,2.3642437E+001] C3 [2.42510891E+001,5.36970367E+001,6.88145638E+000] C4 [8.13146515E+001,2.88190498E+001,4.09245186E+001] C5 [6.70646744E+001,2.53395119E+001,6.14522667E+001] C6 [5.60192604E+001,8.31662598E+001,2.93592377E+001] C1-1 [6.4469986E+001,5.25044632E+001,5.22250557E+001] C1-2 [6.31295433E+001,5.21443062E+001,5.02242126E+001] C1-3 [6.25154915E+001,5.25730362E+001,5.2617794E+001] C1-4 [6.3491375E+001,5.21440697E+001,5.06722069E+001] C1-5 [6.21516266E+001,5.32161064E+001,5.23233032E+001] C1-6 [6.2913269E+001,5.16970291E+001,5.16683655E+001] C1-7 [6.22267456E+001,5.40408363E+001,5.0272541E+001] C1-8 [6.1414093E+001,5.28870888E+001,5.27458E+001] NAME V ________ ____________________________________________________ C1-9 [6.29652252E+001,5.32767754E+001,5.27030106E+001] C1-10 [6.35940704E+001,5.27265244E+001,5.23180656E+001] C1-11 [6.34133224E+001,5.39401283E+001,5.29368248E+001] C1-12 [6.18856697E+001,5.31113129E+001,5.18861504E+001] C1-13 [6.32378883E+001,5.30308647E+001,5.04571724E+001] C1-14 [6.18148689E+001,5.33705482E+001,5.29123802E+001] C1-15 [6.43224258E+001,5.23124084E+001,5.21299057E+001] C2-1 [5.59053535E+001,5.20054626E+001,2.28595486E+001] C2-2 [5.71644516E+001,5.13243408E+001,2.31167526E+001] C2-3 [5.66626244E+001,5.00615959E+001,2.27138176E+001] C2-4 [5.73383865E+001,5.04509125E+001,2.36539135E+001] C2-5 [5.6621357E+001,5.01576767E+001,2.38867531E+001] C2-6 [5.59768562E+001,5.17590942E+001,2.49088764E+001] C2-7 [5.64437904E+001,4.71531525E+001,2.23245487E+001] NAME V ________ ____________________________________________________ C2-8 [5.81449051E+001,5.09049644E+001,2.29072056E+001] C2-9 [5.37190132E+001,4.87386665E+001,2.28188381E+001] C2-10 [5.77416382E+001,4.93461685E+001,2.32014389E+001] C2-11 [5.68353958E+001,5.11093979E+001,2.43693123E+001] C2-12 [5.79631157E+001,5.0297657E+001,2.28039799E+001] C2-13 [5.57930183E+001,5.11965866E+001,2.35887661E+001] C2-14 [5.57345848E+001,5.03228951E+001,2.30780907E+001] C2-15 [5.69435997E+001,4.8590435E+001,2.58747597E+001] C3-1 [2.40239315E+001,5.2352993E+001,5.63517284E+000] C3-2 [2.39717846E+001,5.30635986E+001,5.86633539E+000] C3-3 [2.70314407E+001,5.48788643E+001,7.96345377E+000] C3-4 [2.39875908E+001,5.39634552E+001,5.87654877E+000] C3-5 [2.47772026E+001,5.2187336E+001,6.83652115E+000] C3-6 [2.32920208E+001,5.41494293E+001,6.40737772E+000] NAME V ________ ____________________________________________________ C3-7 [2.46129742E+001,5.32308769E+001,6.29999685E+000] C3-8 [2.51000671E+001,5.33271561E+001,8.86797047E+000] C3-9 [2.4337059E+001,5.26281281E+001,6.9616766E+000] C3-10 [2.39770508E+001,5.42386856E+001,5.63018417E+000] C3-11 [2.59837551E+001,5.34013176E+001,6.97773361E+000] C3-12 [2.40400314E+001,5.25649719E+001,7.2636981E+000] C3-13 [2.13184013E+001,5.28633308E+001,8.3834734E+000] C3-14 [2.50075855E+001,5.21548729E+001,6.88196087E+000] C3-15 [2.53695087E+001,5.60495186E+001,6.76059389E+000] C4-1 [8.28819885E+001,2.95163822E+001,4.03809738E+001] C4-2 [8.18269348E+001,2.95735188E+001,3.99435768E+001] C4-3 [8.2709259E+001,2.90755043E+001,4.07345886E+001] C4-4 [8.18622665E+001,2.88013916E+001,4.1822567E+001] C4-5 [7.99165421E+001,2.89941139E+001,4.09653854E+001] NAME V ________ ____________________________________________________ C4-6 [8.12936707E+001,2.98655643E+001,4.00380211E+001] C4-7 [8.21705704E+001,2.90163479E+001,3.94858704E+001] C4-8 [8.20081329E+001,2.89751148E+001,4.1045887E+001] C4-9 [8.25486298E+001,2.84143009E+001,4.15654945E+001] C4-10 [8.22034149E+001,2.92223415E+001,4.20033302E+001] C4-11 [8.2048996E+001,2.98751106E+001,4.09612732E+001] C4-12 [8.09316025E+001,2.7799057E+001,4.12611198E+001] C4-13 [8.04624023E+001,2.88711109E+001,4.07331085E+001] C4-14 [8.13773575E+001,2.97510109E+001,4.09169846E+001] C4-15 [8.35310364E+001,2.971031E+001,4.16878052E+001] C5-1 [6.87114258E+001,2.53504581E+001,6.11055298E+001] C5-2 [6.73569031E+001,2.35163498E+001,6.01617432E+001] C5-3 [6.78224869E+001,2.61236534E+001,6.0729248E+001] C5-4 [6.76432266E+001,2.56426888E+001,6.35400085E+001] NAME V ________ ____________________________________________________ C5-5 [6.75377045E+001,2.60873699E+001,6.35584145E+001] C5-6 [6.84944687E+001,2.51576214E+001,6.24934502E+001] C5-7 [6.79246216E+001,2.53722992E+001,6.32098122E+001] C5-8 [6.84075165E+001,2.63778133E+001,6.10950584E+001] C5-9 [6.73214798E+001,2.70551453E+001,6.27835197E+001] C5-10 [6.50006485E+001,2.67408028E+001,6.07828026E+001] C5-11 [6.68869705E+001,2.3982399E+001,6.13440819E+001] C5-12 [6.55524521E+001,2.42231808E+001,6.07235756E+001] C5-13 [6.72140808E+001,2.42842178E+001,6.21546478E+001] C5-14 [6.89587936E+001,2.67715569E+001,6.08621559E+001] C5-15 [6.68405685E+001,2.44039059E+001,6.12652893E+001] C6-1 [5.4925251E+001,8.28179474E+001,3.0869236E+001] C6-2 [5.52922363E+001,8.23375549E+001,2.94804363E+001] C6-3 [5.60466652E+001,8.18454132E+001,2.99774895E+001] NAME V ________ ____________________________________________________ C6-4 [5.74460373E+001,8.26830368E+001,2.86887722E+001] C6-5 [5.57439041E+001,8.14622726E+001,2.94924259E+001] C6-6 [5.4913372E+001,8.48766251E+001,2.92711105E+001] C6-7 [5.66876144E+001,8.25907898E+001,2.84199276E+001] C6-8 [5.6253479E+001,8.3280838E+001,2.69524212E+001] C6-9 [5.50792351E+001,8.37676392E+001,3.08755417E+001] C6-10 [5.57719955E+001,8.11036758E+001,2.92569256E+001] C6-11 [5.60834808E+001,8.3103096E+001,3.09748001E+001] C6-12 [5.58962059E+001,8.3612648E+001,2.95026093E+001] C6-13 [5.73348083E+001,8.26950226E+001,2.88242455E+001] C6-14 [5.45099411E+001,8.33315659E+001,2.90559101E+001] C6-15 [5.57930641E+001,8.5720871E+001,2.92863998E+001] C1-97 [6.3716671E+001,5.41518326E+001,5.18371048E+001] C1-98 [6.58774261E+001,5.32223206E+001,5.05089798E+001] NAME V _________ ____________________________________________________ C1-99 [6.31867676E+001,5.25712204E+001,5.16621819E+001] C1-100 [6.20503845E+001,5.15550919E+001,5.08155479E+001] 100 rows selected.To visualize the resulting vectors in space, consider the following graph:
- Run a similarity search on the generated vectors in the
genvectable.First define a variable called
cluster_numberto be used to form the name of the vector in the query.DEFINE cluster_number = '&clusterid'You will be prompted to enter a value for
clusterid. In this example, we use5:Enter value for clusterid: 5Run the following query to perform a similarity search on the generated vectors:
SELECT name FROM genvec ORDER BY VECTOR_DISTANCE(v,(SELECT v FROM genvec WHERE name='C'||'&cluster_number'),EUCLIDEAN) FETCH EXACT FIRST 20 ROWS ONLY;Example output:
NAME ________ C5 C5-15 C5-13 C5-3 C5-11 C5-1 C5-8 C5-6 C5-7 C5-12 C5-9 C5-4 C5-2 C5-5 NAME ________ C5-14 C5-10 C4-5 C4-12 C4-4 C4-13 20 rows selected. - Here is another example of running a similarity search on the generated vectors, this
time using the Cosine distance metric.
SELECT name FROM genvec ORDER BY VECTOR_DISTANCE(v,(SELECT v FROM genvec WHERE name='C'||'&cluster_number'),COSINE) FETCH EXACT FIRST 20 ROWS ONLY;Example output:
NAME ________ C5 C5-6 C5-12 C5-15 C5-7 C5-4 C5-5 C5-13 C5-11 C5-3 C5-1 C5-8 C5-9 C5-2 NAME ________ C5-14 C5-10 C4-5 C4-4 C4-10 C4-12 20 rows selected. - Create a variable called
query_vectorand then useSELECT INTOto store a vector value in the variable.VARIABLE query_vector CLOB BEGIN SELECT v INTO :query_vector FROM genvec WHERE name='C'||'&cluster_number'; END; / PRINT query_vector;Example output:
QUERY_VECTOR -------------------------------------------------------------------- [6.70646744E+001,2.53395119E+001,6.14522667E+001] - Create an explain plan for a similarity search using the query vector created in the
previous step.
EXPLAIN PLAN FOR SELECT name FROM genvec ORDER BY VECTOR_DISTANCE(v, :query_vector, EUCLIDEAN) FETCH EXACT FIRST 20 ROWS ONLY; SELECT plan_table_output FROM table(dbms_xplan.display('plan_table',null,'all'));Example output:PLAN_TABLE_OUTPUT _____________________________________________________________________________________ Plan hash value: 1549136425 ---------------------------------------------------------------------------------- | Id | Operation | Name | Rows | Bytes | Cost (%CPU)| Time | ---------------------------------------------------------------------------------- | 0 | SELECT STATEMENT | | 20 | 5040 | 4 (25)| 00:00:01 | |* 1 | COUNT STOPKEY | | | | | | | 2 | VIEW | | 100 | 25200 | 4 (25)| 00:00:01 | |* 3 | SORT ORDER BY STOPKEY| | 100 | 25800 | 4 (25)| 00:00:01 | | 4 | TABLE ACCESS FULL | GENVEC | 100 | 25800 | 3 (0)| 00:00:01 | ---------------------------------------------------------------------------------- Query Block Name / Object Alias (identified by operation id): ------------------------------------------------------------- PLAN_TABLE_OUTPUT ______________________________________________________________ 1 - SEL$2 2 - SEL$1 / "from$_subquery$_002"@"SEL$2" 3 - SEL$1 4 - SEL$1 / "GENVEC"@"SEL$1" Predicate Information (identified by operation id): --------------------------------------------------- 1 - filter(ROWNUM<=20) 3 - filter(ROWNUM<=20) Column Projection Information (identified by operation id): ----------------------------------------------------------- PLAN_TABLE_OUTPUT __________________________________________________________________________ 1 - "from$_subquery$_002"."NAME"[VARCHAR2,500] 2 - "from$_subquery$_002"."NAME"[VARCHAR2,500] 3 - (#keys=1) VECTOR_DISTANCE("V" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[BINARY_DOUBLE,8], "NAME"[VARCHAR2,500] 4 - "NAME"[VARCHAR2,500], VECTOR_DISTANCE("V" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[BINARY_DOUBLE,8] Note ----- - dynamic statistics used: dynamic sampling (level=2) 41 rows selected. - Create an Hierarchical Navigable Small World (HNSW) index.
CREATE VECTOR INDEX genvec_hnsw_idx ON genvec(v) ORGANIZATION INMEMORY NEIGHBOR GRAPH DISTANCE EUCLIDEAN WITH TARGET ACCURACY 95; SELECT INDEX_NAME, INDEX_TYPE, INDEX_SUBTYPE FROM USER_INDEXES;Example output:
INDEX_NAME INDEX_TYPE INDEX_SUBTYPE ___________________________ _____________ _______________________________ DM$CEDOC_MODEL NORMAL SYS_IL0000073592C00002$$ LOB GENVEC_HNSW_IDX VECTOR INMEMORY_NEIGHBOR_GRAPH_HNSW SYS_C008694 NORMAL 4 rows selected. - Query information about the HNSW index from the
VECSYS.VECTOR$INDEXview.SELECT JSON_SERIALIZE(IDX_PARAMS RETURNING VARCHAR2 PRETTY) FROM VECSYS.VECTOR$INDEX WHERE IDX_NAME = 'GENVEC_HNSW_IDX';Example output:
JSON_SERIALIZE(IDX_PARAMSRETURNINGVARCHAR2PRETTY) ______________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________ { "type" : "HNSW", "num_neighbors" : 32, "efConstruction" : 300, "distance" : "EUCLIDEAN", "accuracy" : 95, "vector_type" : "FLOAT32", "vector_dimension" : 3, "degree_of_parallelism" : 1, "pdb_id" : 3, "indexed_col" : "V" }For information about the columns of the
VECSYS.VECTOR$INDEXview, see VECSYS.VECTOR$INDEX. - With the HNSW index created, create another explain plan for a similarity search on the
genvectable.EXPLAIN PLAN FOR SELECT name FROM genvec ORDER BY VECTOR_DISTANCE(v, :query_vector, EUCLIDEAN) FETCH APPROX FIRST 20 rows only; SELECT plan_table_output FROM table(dbms_xplan.display('plan_table',null,'all'));Example output:
PLAN_TABLE_OUTPUT _____________________________________________________________________________________________________________ Plan hash value: 1202819565 ---------------------------------------------------------------------------------------------------------- | Id | Operation | Name | Rows | Bytes |TempSpc| Cost (%CPU)| Time | ---------------------------------------------------------------------------------------------------------- | 0 | SELECT STATEMENT | | 20 | 5040 | | 165 (2)| 00:00:01 | |* 1 | COUNT STOPKEY | | | | | | | | 2 | VIEW | | 100 | 25200 | | 165 (2)| 00:00:01 | |* 3 | SORT ORDER BY STOPKEY | | 100 | 425K| 808K| 165 (2)| 00:00:01 | | 4 | TABLE ACCESS BY INDEX ROWID| GENVEC | 100 | 425K| | 1 (0)| 00:00:01 | | 5 | VECTOR INDEX HNSW SCAN | GENVEC_HNSW_IDX | 100 | 425K| | 1 (0)| 00:00:01 | ---------------------------------------------------------------------------------------------------------- Query Block Name / Object Alias (identified by operation id): PLAN_TABLE_OUTPUT ________________________________________________________________ ------------------------------------------------------------- 1 - SEL$2 2 - SEL$1 / "from$_subquery$_002"@"SEL$2" 3 - SEL$1 4 - SEL$1 / "GENVEC"@"SEL$1" 5 - SEL$1 / "GENVEC"@"SEL$1" Predicate Information (identified by operation id): --------------------------------------------------- 1 - filter(ROWNUM<=20) 3 - filter(ROWNUM<=20) PLAN_TABLE_OUTPUT _____________________________________________________________________________________________________ Column Projection Information (identified by operation id): ----------------------------------------------------------- 1 - "from$_subquery$_002"."NAME"[VARCHAR2,500] 2 - "from$_subquery$_002"."NAME"[VARCHAR2,500] 3 - (#keys=1) VECTOR_DISTANCE("V" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[8], "NAME"[VARCHAR2,500] 4 - "NAME"[VARCHAR2,500], VECTOR_DISTANCE("V" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[8] 5 - "GENVEC".ROWID[ROWID,10], VECTOR_DISTANCE("V" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[8] Note ----- PLAN_TABLE_OUTPUT ___________________________________________________________ - dynamic statistics used: dynamic sampling (level=2) 43 rows selected.As you can see, the explain plan now includes information about the HNSW index.
- Again perform a similarity search on the vectors in the
genvectable. Note that it is possible for query results to vary based on the indexing technique used. The results included in this scenario are simply an example.SELECT name FROM genvec ORDER BY vector_distance(v, :query_vector, EUCLIDEAN) FETCH APPROX FIRST 20 ROWS ONLY;Example output:
NAME ________ C5 C5-15 C5-13 C5-3 C5-11 C5-1 C5-8 C5-6 C5-7 C5-12 C5-9 C5-4 C5-2 C5-5 NAME ________ C5-14 C5-10 C4-5 C4-12 C4-4 C4-13 20 rows selected. - Drop the HNSW index and create an Inverted File Flat (IVF) vector index.
DROP INDEX genvec_hnsw_idx; CREATE VECTOR INDEX genvec_ivf_idx ON genvec(v) ORGANIZATION NEIGHBOR PARTITIONS DISTANCE EUCLIDEAN WITH TARGET ACCURACY 95; SELECT JSON_SERIALIZE(IDX_PARAMS RETURNING VARCHAR2 PRETTY) FROM VECSYS.VECTOR$INDEX WHERE IDX_NAME = 'GENVEC_IVF_IDX';Example output:
JSON_SERIALIZE(IDX_PARAMSRETURNINGVARCHAR2PRETTY) ________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________________ { "target_centroids" : 40, "pdb_id" : 3, "vector_type" : "FLOAT32", "type" : "IVF_FLAT", "vector_dimension" : 3, "distance" : "EUCLIDEAN", "indexed_col" : "V", "min_vectors_per_partition" : 10, "degree_of_parallelism" : 1, "accuracy" : 95, "num_centroids" : 24, "samples_per_partition" : 256 } - Again create an explain plan, which now includes information about the newly created
IVF index.
EXPLAIN PLAN FOR SELECT name FROM genvec ORDER BY VECTOR_DISTANCE(v, :query_vector, EUCLIDEAN) FETCH APPROX FIRST 20 ROWS ONLY; SELECT plan_table_output FROM table(dbms_xplan.display('plan_table',null,'all'));Example output:
PLAN_TABLE_OUTPUT __________________________________________________________________________________________________________________________________________________________________________ Plan hash value: 2965029064 ----------------------------------------------------------------------------------------------------------------------------------------------------------------------- | Id | Operation | Name | Rows | Bytes | Cost (%CPU)| Time | Pstart| Pstop | ----------------------------------------------------------------------------------------------------------------------------------------------------------------------- | 0 | SELECT STATEMENT | | 5 | 1260 | 29 (14)| 00:00:01 | | | | 1 | VIEW | | 5 | 1260 | 29 (14)| 00:00:01 | | | | 2 | SORT ORDER BY | | 5 | 21910 | 29 (14)| 00:00:01 | | | |* 3 | HASH JOIN | | 5 | 21910 | 28 (11)| 00:00:01 | | | | 4 | VIEW | VW_IVPSR_11E7D7DE | 20 | 320 | 24 (9)| 00:00:01 | | | |* 5 | COUNT STOPKEY | | | | | | | | | 6 | VIEW | VW_IVPSJ_578B79F1 | 25 | 450 | 24 (9)| 00:00:01 | | | |* 7 | SORT ORDER BY STOPKEY | | 25 | 550 | 24 (9)| 00:00:01 | | | |* 8 | HASH JOIN | | 25 | 550 | 23 (5)| 00:00:01 | | | PLAN_TABLE_OUTPUT __________________________________________________________________________________________________________________________________________________________________________ | 9 | PART JOIN FILTER CREATE | :BF0000 | 6 | 18 | 4 (25)| 00:00:01 | | | | 10 | VIEW | VW_IVCR_B5B87E67 | 6 | 18 | 4 (25)| 00:00:01 | | | |* 11 | COUNT STOPKEY | | | | | | | | | 12 | VIEW | VW_IVCN_9A1D2119 | 24 | 312 | 4 (25)| 00:00:01 | | | |* 13 | SORT ORDER BY STOPKEY | | 24 | 216 | 4 (25)| 00:00:01 | | | | 14 | TABLE ACCESS FULL | VECTOR$GENVEC_IVF_IDX$87355_87370_0$IVF_FLAT_CENTROIDS | 24 | 216 | 3 (0)| 00:00:01 | | | | 15 | PARTITION LIST JOIN-FILTER| | 100 | 1900 | 3 (0)| 00:00:01 |:BF0000|:BF0000| | 16 | TABLE ACCESS FULL | VECTOR$GENVEC_IVF_IDX$87355_87370_0$IVF_FLAT_CENTROID_PARTITIONS | 100 | 1900 | 3 (0)| 00:00:01 |:BF0000|:BF0000| | 17 | TABLE ACCESS FULL | GENVEC | 100 | 426K| 3 (0)| 00:00:01 | | | ----------------------------------------------------------------------------------------------------------------------------------------------------------------------- Query Block Name / Object Alias (identified by operation id): ------------------------------------------------------------- PLAN_TABLE_OUTPUT ___________________________________________________________ 1 - SEL$94C0F189 / "from$_subquery$_002"@"SEL$2" 2 - SEL$94C0F189 4 - SEL$E731354C / "VW_IVPSR_11E7D7DE"@"SEL$1" 5 - SEL$E731354C 6 - SEL$0C00A749 / "VW_IVPSJ_578B79F1"@"SEL$E731354C" 7 - SEL$0C00A749 10 - SEL$700CE8F1 / "VW_IVCR_B5B87E67"@"SEL$0C00A749" 11 - SEL$700CE8F1 12 - SEL$E5326247 / "VW_IVCN_9A1D2119"@"SEL$700CE8F1" 13 - SEL$E5326247 14 - SEL$E5326247 / "VTIX_CENTRD"@"SEL$E5326247" 16 - SEL$0C00A749 / "VTIX_CNPART"@"SEL$0C00A749" 17 - SEL$94C0F189 / "GENVEC"@"SEL$1" PLAN_TABLE_OUTPUT ______________________________________________________________________________ Predicate Information (identified by operation id): --------------------------------------------------- 3 - access("GENVEC".ROWID="VW_IVPSR_11E7D7DE"."BASE_TABLE_ROWID") 5 - filter(ROWNUM<=20) 7 - filter(ROWNUM<=20) 8 - access("VW_IVCR_B5B87E67"."CENTROID_ID"="VTIX_CNPART"."CENTROID_ID") 11 - filter(ROWNUM<=6) 13 - filter(ROWNUM<=6) Column Projection Information (identified by operation id): ----------------------------------------------------------- 1 - "from$_subquery$_002"."NAME"[VARCHAR2,500] PLAN_TABLE_OUTPUT __________________________________________________________________________________________________________________________________________________________________ 2 - (#keys=1) "VEC_DIST"[BINARY_DOUBLE,8], "GENVEC"."NAME"[VARCHAR2,500] 3 - (#keys=1) "GENVEC"."NAME"[VARCHAR2,500], "VEC_DIST"[BINARY_DOUBLE,8], "GENVEC"."NAME"[VARCHAR2,500] 4 - "BASE_TABLE_ROWID"[ROWID,10], "VEC_DIST"[BINARY_DOUBLE,8] 5 - "VW_IVPSJ_578B79F1"."BASE_TABLE_ROWID"[ROWID,10], "VW_IVPSJ_578B79F1"."VEC_DIST"[BINARY_DOUBLE,8] 6 - "VW_IVPSJ_578B79F1"."BASE_TABLE_ROWID"[ROWID,10], "VW_IVPSJ_578B79F1"."VEC_DIST"[BINARY_DOUBLE,8] 7 - (#keys=1) VECTOR_DISTANCE("VTIX_CNPART"."DATA_VECTOR" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[BINARY_DOUBLE,8], "VTIX_CNPART"."BASE_TABLE_ROWID"[ROWID,10] 8 - (#keys=1) "VTIX_CNPART"."BASE_TABLE_ROWID"[ROWID,10], VECTOR_DISTANCE("VTIX_CNPART"."DATA_VECTOR" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[BINARY_DOUBLE,8] 9 - "VW_IVCR_B5B87E67"."CENTROID_ID"[NUMBER,22], "VW_IVCR_B5B87E67"."CENTROID_ID"[NUMBER,22] 10 - "CENTROID_ID"[NUMBER,22] 11 - "VW_IVCN_9A1D2119"."CENTROID_ID"[NUMBER,22] 12 - "VW_IVCN_9A1D2119"."CENTROID_ID"[NUMBER,22] 13 - (#keys=1) VECTOR_DISTANCE("VECTOR$GENVEC_IVF_IDX$87355_87370_0$IVF_FLAT_CENTROIDS"."CENTROID_VECTOR" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * PLAN_TABLE_OUTPUT _________________________________________________________________________________________________________________________________________________________________ /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[BINARY_DOUBLE,8], "VTIX_CENTRD"."CENTROID_ID"[NUMBER,22] 14 - "VTIX_CENTRD"."CENTROID_ID"[NUMBER,22], VECTOR_DISTANCE("VECTOR$GENVEC_IVF_IDX$87355_87370_0$IVF_FLAT_CENTROIDS"."CENTROID_VECTOR" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[BINARY_DOUBLE,8] 15 - "VTIX_CNPART"."BASE_TABLE_ROWID"[ROWID,10], "VTIX_CNPART"."CENTROID_ID"[NUMBER,22], VECTOR_DISTANCE("VTIX_CNPART"."DATA_VECTOR" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[BINARY_DOUBLE,8] 16 - "VTIX_CNPART"."BASE_TABLE_ROWID"[ROWID,10], "VTIX_CNPART"."CENTROID_ID"[NUMBER,22], VECTOR_DISTANCE("VTIX_CNPART"."DATA_VECTOR" /*+ LOB_BY_VALUE */ , VECTOR(:QUERY_VECTOR, *, * /*+ USEBLOBPCW_QVCGMD */ ), EUCLIDEAN)[BINARY_DOUBLE,8] 17 - "GENVEC".ROWID[ROWID,10], "GENVEC"."NAME"[VARCHAR2,500] Note ----- - dynamic statistics used: dynamic sampling (level=2) - this is an adaptive plan 83 rows selected. - Finally, run the similarity search once again.
SELECT name FROM genvec ORDER BY VECTOR_DISTANCE(v, :query_vector, EUCLIDEAN) FETCH APPROX FIRST 20 ROWS ONLY;Example output:
NAME ________ C5 C5-15 C5-13 C5-3 C5-11 C5-1 C5-8 C5-6 C5-7 C5-12 C5-9 C5-4 C5-2 C5-5 NAME ________ C5-14 C5-10 C4-5 C4-12 C4-4 C4-13 20 rows selected.
Parent topic: Get Started